Identification and Pathogenicity Analysis of a Novel Fibrinogen Bβ Chain p.Gly293Val Variant Causing Hypofibrinogenemia
DOI:
https://doi.org/10.14740/jh2183Keywords:
Hypofibrinogenemia, Fibrinogen, FGB, VariantAbstract
Background: Hypofibrinogenemia is a rare bleeding disorder characterized by excessive bleeding, impaired wound healing, and elevated perioperative risk. It most commonly results from pathogenic variants in the FGB gene. This study aimed to analyze the clinical phenotypes and genetic variants in a family with hypofibrinogenemia and explore its molecular pathogenic mechanisms.
Methods: Fibrinogen (Fg) activity (Fg:C) was measured using the Clauss method and the prothrombin time (PT)-derived method, and Fg antigen (Fg:Ag) levels were determined by enzyme-linked immunosorbent assay (ELISA). Fg polymerization capacity was evaluated via a thrombin-induced Fg polymerization assay, and Fg levels and function were assessed using thromboelastography. Sanger sequencing was performed to screen for variants in all exons and flanking regions of the FGA, FGB, and FGG genes. Multiple in silico tools, including ClustalX-2.1-win, MutationTaster, PolyPhen-2, PROVEAN, I-Mutant 2.0 and Swiss-Pdb Viewer, were used to assess the conservation of the variation sites and their impact on protein structure and function. The pathogenicity of the variation sites was evaluated according to the American College of Medical Genetics and Genomics (ACMG) standards and guidelines for the interpretation of sequence variants.
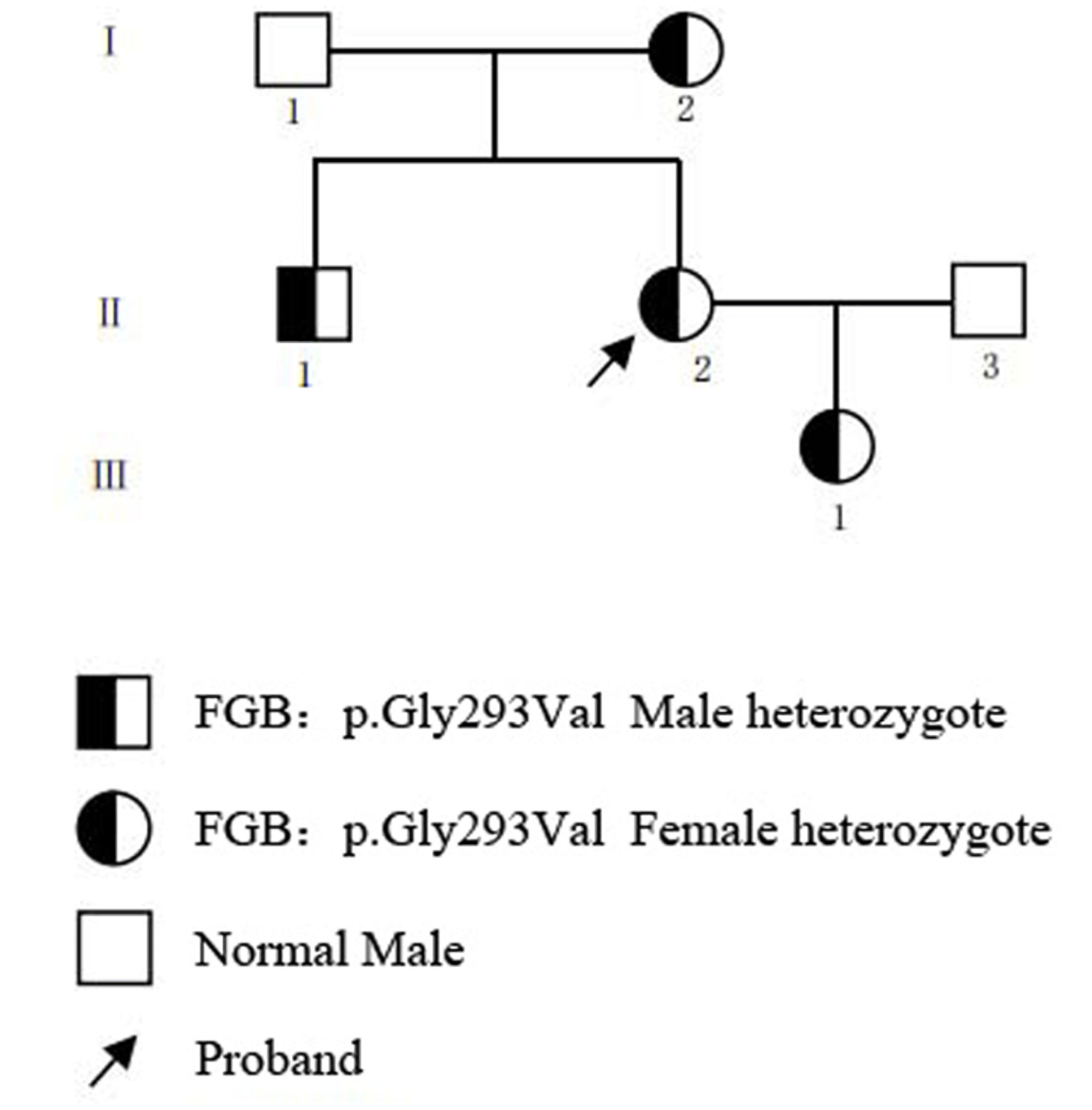
Results: The proband and affected members exhibited prolonged thrombin time (TT), reduced Fg:C and Fg:Ag levels, and a hypocoagulable thromboelastography profile. Notably, Fg polymerization kinetics remained preserved, consistent with hypofibrinogenemia rather than dysfibrinogenemia. Genetic analysis identified a heterozygous missense variant FGB c.878G>T (p.Gly293Val) segregating with the phenotype. This variant was absent from population databases, located at a highly conserved residue, and predicted to be deleterious by multiple in silico tools. Protein structural modeling indicated local conformational disturbance. The variant was classified as likely pathogenic following the 2015 ACMG/Association for Molecular Pathology (AMP) standard guidelines.
Conclusions: The FGB p.Gly293Val variant may cause a significant decrease in Fg:C and Fg:Ag by disrupting the structure and function of the Fg protein.
Published
Issue
Section
License
Copyright (c) 2026 The authors

This work is licensed under a Creative Commons Attribution 4.0 International License.








